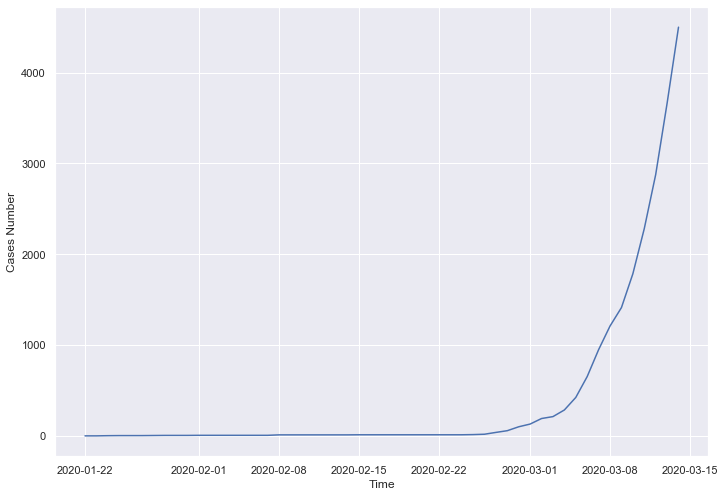
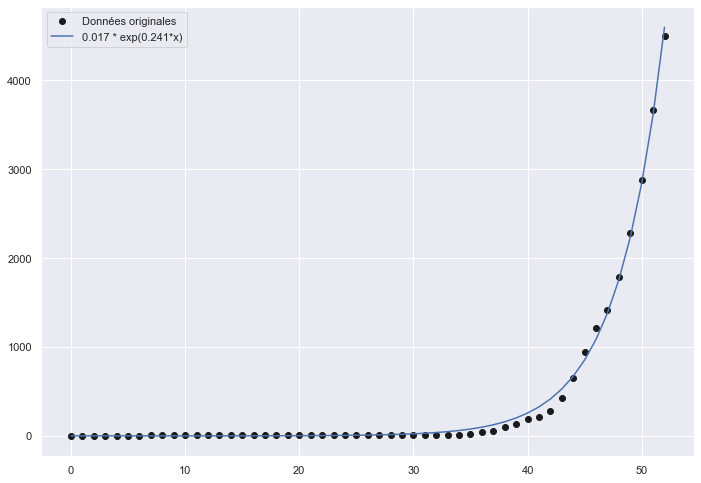
- J’ai installé une nouvelle lib avec pip install
et maintenant tout est cassé - J’ai un venv que je veux reproduire en prod, comment faire ?
- J’ai installé python 3.10.2 et 3.11.2 mais je ne comprend pas comment utiliser l’un ou l’autre ?
- C’est quoi ce tas de trucs incomprehensibles avec pip, easy_install, poetry, conda, virtualenv, pdm ?
- Pourquoi des fois je vois des setup.py, et parfois des pyproject.toml ? Pourquoi des lockfiles pdm.lock ou poetry.lock ?
- J’ai mon venv qui marche niquel, avec pleins de belles librairies de data science, je fais un pip install d’une nouvelle lib et paf, tout est cassé, ça marche pas je ne sais pas pourquoi ?
- JE VEUX JUSTE UN TRUC SIMPLE QUI ME PERMETTE DE DEV DANS DES NOTEBOOKS DE MANIERE PRAGMATIQUE COMMENT JE FAIS ?
Pré-requis:
Avoir soit un linux sur lequel on peut installer ce qu’on veut, soit un wsl sous windows pour pouvoir travailler efficacement (l’install est simple désormais: https://learn.microsoft.com/en-us/windows/wsl/install), avoir un python d’installé (il y en a normalement un par défaut dans la plupart des linux couramment utilisés), ainsi que pip (idem)
J’ai installé une nouvelle lib avec pip install pandas et maintenant tout est cassé
Lorsqu’on fait un:
pip install pandas
directement dans le terminal sans autre précaution, on installe une librairie dans le système global, ça pose problème:
- Si on a un autre projet qui a besoin d’une autre version de pandas que ce projet courant, on ne pourra pas faire co-exister les deux
- On risque de modifier la version de pandas utilisée par l’autre projet (et donc le casser)
Dans ces deux cas on peut avoir des messages d’erreur parfois un peu obscurs, nous disant qu’il y a conflit.
Pour résoudre ce problème, en python on utilise des “environnements virtuels”, c’est à dire un mécanisme qui permet d’isoler les version de python et les dépendances.
Généralement, on va vouloir avoir un environnement virtuel (ou venv) par projet.
Pour en créer un:
python -m venv projet_a_env
Puis on l’active (on se “met dedans”)
source projet_a_env/bin/activate
On peut alors simplement installer les librairies qu’on souhaite:
pip install pandas
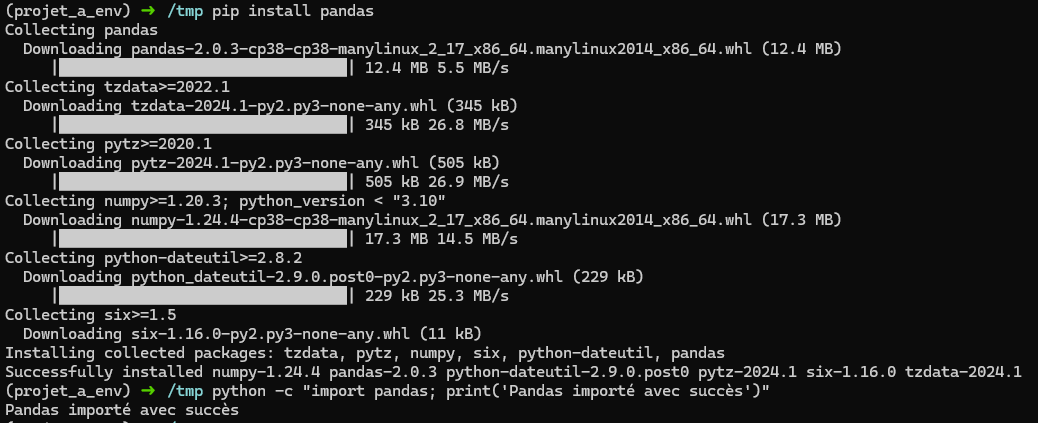
Si on veut sortir de l’environnement virtuel:
deactivate
On revient alors dans le système global, pandas n’est plus installé:
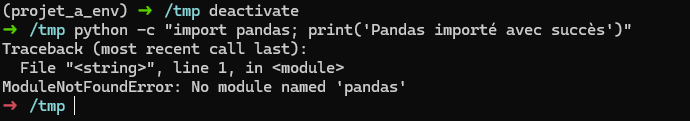
J’ai un venv que je veux reproduire en prod, comment faire ?
pip freeze > requirements.txt
Le fichier requirements.txt est ici de la forme:
numpy==1.24.4
pandas==2.0.3
python-dateutil==2.9.0.post0
pytz==2024.1
six==1.16.0
tzdata==2024.1
Puis une fois dans le nouveau venv de prod :
pip install -r requirements.txt
J’ai installé python 3.10.2 et 3.11.2 mais je ne comprend pas comment utiliser l’un ou l’autre ?
On peut vite se retrouver à devoir gérer plusieurs versions de python sur la même machine (dépendamment des projets sur lesquels on travaille). Pour gérer ce problème, le plus simple est d’installer et d’utiliser pyenv:
curl https://pyenv.run | bash
puis:
echo 'export PYENV_ROOT="$HOME/.pyenv"' >> ~/.bashrc
echo 'command -v pyenv >/dev/null || export PATH="$PYENV_ROOT/bin:$PATH"' >> ~/.bashrc
echo 'eval "$(pyenv init -)"' >> ~/.bashrc
(si vous utilisez zsh ou un autre shell, voir la doc https://github.com/pyenv/pyenv)
Ensuite on va installer les dépendances nécessaires (ici sur ubuntu/debian) pour pouvoir compiler d’autres versions de python:
sudo apt update
sudo apt install -y make build-essential libssl-dev zlib1g-dev \
libbz2-dev libreadline-dev libsqlite3-dev wget curl llvm \
libncursesw5-dev xz-utils tk-dev libxml2-dev libxmlsec1-dev libffi-dev liblzma-dev
Ensuite, c’est simple, pour installer une version de python sur le système:
pyenv install 3.10.2
Puis pour switcher vers une version de python spécifiquement pour le projet courant:
pyenv local 3.10.2
Ceci créé un fichier .python-version, qui contient donc la version de python du projet.
Quand vous créérez un venv, il sera automatiquement utilisé.
C’est quoi ce tas de trucs incomprehensibles avec pip, easy_install, poetry, conda, virtualenv, pdm ?
- pip: gestionnaire de paquets standard de python
- easy_install: un ancien outil d’installation de paquet, une autre époque, oublie
- poetry: un gestionnaire de paquet plus moderne qui résout pas mal de problemes de pip, notamment la gestion de conflits de paquets dans un venv, ainsi que la gestion de dépendances primaires et secondaires (voir après).
- conda: Gestionnaire de paquet et d’environnements pythons, populaire dans la communauté scientifique. J’ai eu plusieurs soucis d’exports windows/linux pour reproduire des environnements condas, ne suit pas vraiment les standards python, utilise son propre mécanisme d’environnements virtuels. Peut être bien dans un cadre scientifique, mais pas dans les cadres industriels que j’ai rencontrés.
- pdm: Comme poetry, en mieux: respecte les standards python, permet d’avoir plus de contrôle sur la construction et le publish des wheels, notamment.
Pourquoi des fois je vois des setup.py, et parfois des pyproject.toml ? Pourquoi des lockfiles pdm.lock ou poetry.lock ?
- setup.py: permet d’installer le projet courant comme paquet (pip l’utilisait quand on faisait pip install
). C’est l’ancienne façon de faire, à oublier - pyproject.toml: standard depuis 2016 qui remplace le setup.py, et est le point de configuration central du projet, i.e., quelles sont les dependances primaires, de dev, comment build le projet, comment le distribuer, quelles sont les metadonnées, etc.
Lorsqu’on fait un “pip install pandas” et qu’on liste les dépendances, on a vu plus haut que ça nous donne une liste de toutes les librairies installées, c’est à dire à la fois les dépendances primaires, et les dépendances secondaires, si bien qu’on peut se retrouver perdu à ne plus s’y retrouver. Avec le pyprojet.toml, on obtient une liste de toutes les dépendances primaires:
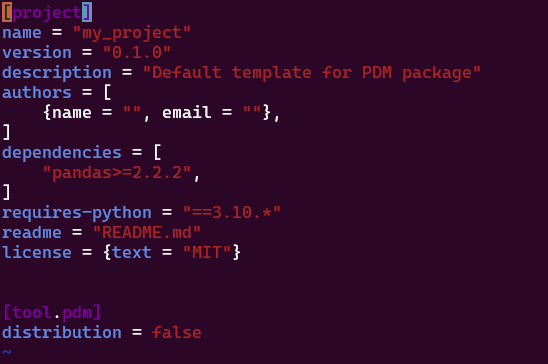
Dans le fichier pdm.lock ou poetry.lock on va avoir les dépendances secondaires, c’est à dire les dépendances de dépendances. Elles vont être utiles pour pouvoir recréer le venv à l’identique ailleurs:
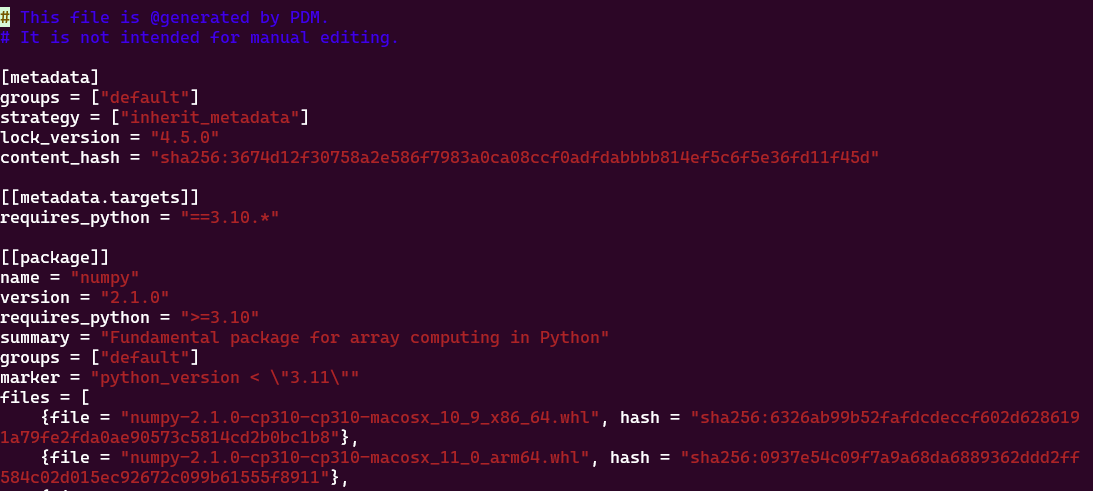
J’ai mon venv qui marche bien, avec pleins de belles librairies pour la data science, je fais un pip install d’une nouvelle lib et paf, tout est cassé, ça marche pas ?
Il y a en fait deux cas:
- Ca casse notre env car il y a une incompatibilité fondamentale et ça met le bazar sans nous dire pourquoi, c’est moche
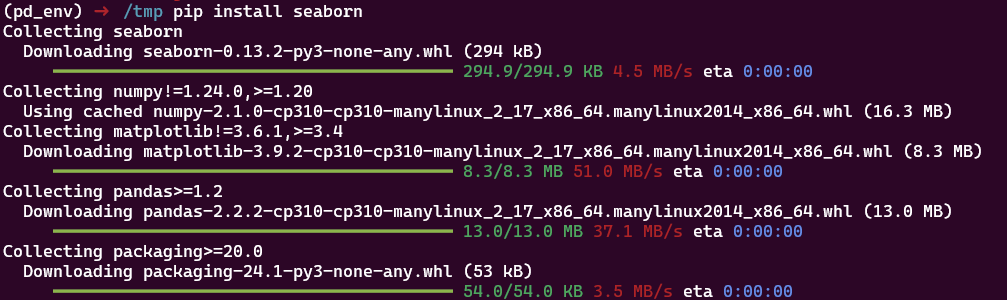
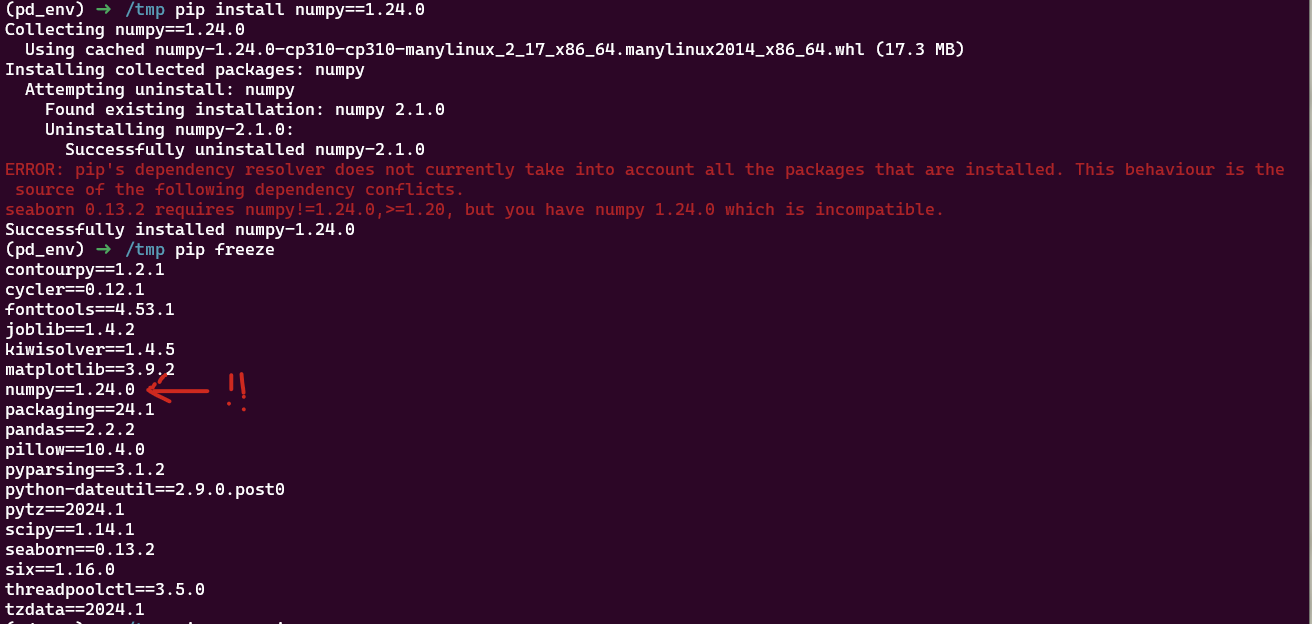
Ici on installe d’abord seaborn puis numpy dans une version incompatibl. Seaborn ne fonctionne plus: on a certes un message d’erreur, mais pas de rollback, le venv est dans état non fonctionnel pour notre code.
- Ca upgrade automatiquement une lib précédente, qu’on avait pourtant fixée :
Par exemple on installe numpy dans une version 1.1.15, on fait notre code, puis plus tard on install seaborn. A ce moment là, numpy va automatiquement être mis à jour vers une version plus récente (sans nous demander notre avis), et donc potentiellement ne sera plus compatible avec notre ancien code.
En fait dans ces deux cas nous sommes dans des exemples de “dependency hell”:
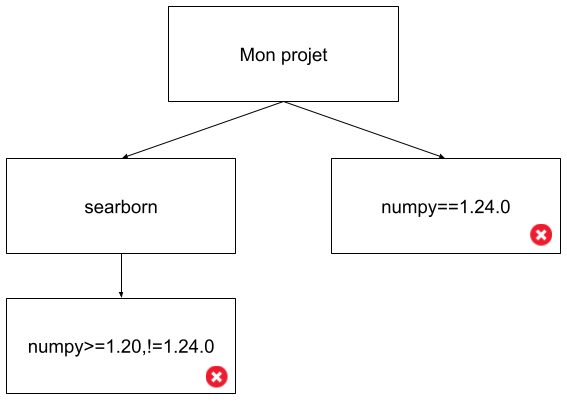
Ici on est dans un cas simple, mais imaginez la complexité lorsqu’on a des dépendances au 5ème degré, avec des dizaines de librairies…
Pour mieux gérer ces problèmes, qui peuvent vite vous faire perdre des années de vie, je conseille d’utiliser pdm (https://pdm-project.org/en/latest/), et ce pour plusieurs raisons:
- Le rollback fonctionne, si on essaie d’installer quelque chose, on revient dans l’état du venv précédent qui lui était fonctionnel
- Lorsqu’on essaie d’installer une librairie où il y a un conflit, on a un message clair qui nous guide pour trouver une solution
- On a une séparation claire des dépendances primaires et secondaires
- On respecte les normes pep strictement (ce qui n’est pas le cas de poetry, par exemple)
- On peut choisir la façon de build ses wheels pour distribuer son programme, contrairement à poetry qui force son propre outil de build (qui a déjà été bloquant pour moi)
- Il est assez efficace
JE VEUX JUSTE UN TRUC SIMPLE QUI ME PERMETTE DE DEV DANS DES NOTEBOOKS DE MANIERE PRAGMATIQUE COMMENT JE FAIS ?
Si on est sous windows, installation du wsl en administrateur, le powershell on oublie:
wsl --install
Installation de pdm:
sudo apt install python3.10-venv
curl -sSL https://pdm-project.org/install-pdm.py | python3 -
echo export PATH=/home/romain/.local/bin:$PATH >> ~/.bashrc
source ~/.bashrc
Installation de pyenv et choix de la version de python:
curl https://pyenv.run | bash
echo 'export PYENV_ROOT="$HOME/.pyenv"' >> ~/.bashrc
echo 'command -v pyenv >/dev/null || export PATH="$PYENV_ROOT/bin:$PATH"' >> ~/.bashrc
echo 'eval "$(pyenv init -)"' >> ~/.bashrc
sudo apt update
sudo apt install -y make build-essential libssl-dev zlib1g-dev \
libbz2-dev libreadline-dev libsqlite3-dev wget curl llvm \
libncursesw5-dev xz-utils tk-dev libxml2-dev libxmlsec1-dev libffi-dev liblzma-dev
pyenv install 3.11.2
pyenv local 3.11.2
Creation du dossier de travail
mkdir my_project
Creation de l’environnement virtuel (répondre aux questions qui apparaissent pour initialiser le projet)
pdm init
Ajout de librairie(s)
pdm add pandas
Ajout de ipykernel pour pouvoir utiliser notre venv directement dans notre notebook
pdm add ipykernel
Attention: pdm créée par defaut le venv dans .venv, faire un “ls - a” pour le voir
On peut alors sélectionner l’environnement virtuel directement dans le notebook, dans vscode par exemple.
]]>